Is An Organism's Genetic Makeup Working Behind The Scenes
Chapter 10: Introduction to Biotechnology
10.1 Cloning and Genetic Engineering
Learning Objectives
By the stop of this section, you lot will be able to:
- Explain the basic techniques used to manipulate genetic cloth
- Explain molecular and reproductive cloning
Biotechnology is the utilize of artificial methods to alter the genetic material of living organisms or cells to produce novel compounds or to perform new functions. Biotechnology has been used for improving livestock and crops since the outset of agriculture through selective convenance. Since the discovery of the structure of Deoxyribonucleic acid in 1953, and particularly since the development of tools and methods to manipulate DNA in the 1970s, biotechnology has become synonymous with the manipulation of organisms' Deoxyribonucleic acid at the molecular level. The main applications of this technology are in medicine (for the production of vaccines and antibiotics) and in agriculture (for the genetic modification of crops). Biotechnology also has many industrial applications, such as fermentation, the treatment of oil spills, and the production of biofuels, too as many household applications such as the use of enzymes in laundry detergent.
Manipulating Genetic Material
To achieve the applications described to a higher place, biotechnologists must be able to extract, manipulate, and clarify nucleic acids.
Review of Nucleic Acid Structure
To understand the basic techniques used to work with nucleic acids, remember that nucleic acids are macromolecules made of nucleotides (a sugar, a phosphate, and a nitrogenous base). The phosphate groups on these molecules each have a net negative charge. An entire set of Deoxyribonucleic acid molecules in the nucleus of eukaryotic organisms is called the genome. Dna has two complementary strands linked by hydrogen bonds between the paired bases.
Unlike DNA in eukaryotic cells, RNA molecules exit the nucleus. Messenger RNA (mRNA) is analyzed nearly frequently because information technology represents the protein-coding genes that are being expressed in the cell.
Isolation of Nucleic Acids
To report or dispense nucleic acids, the Dna must first exist extracted from cells. Various techniques are used to extract different types of Deoxyribonucleic acid (Figure 10.2). Almost nucleic acid extraction techniques involve steps to break open the cell, and then the use of enzymatic reactions to destroy all undesired macromolecules. Cells are broken open using a detergent solution containing buffering compounds. To foreclose degradation and contamination, macromolecules such as proteins and RNA are inactivated using enzymes. The Deoxyribonucleic acid is so brought out of solution using alcohol. The resulting DNA, considering information technology is made up of long polymers, forms a gelatinous mass.
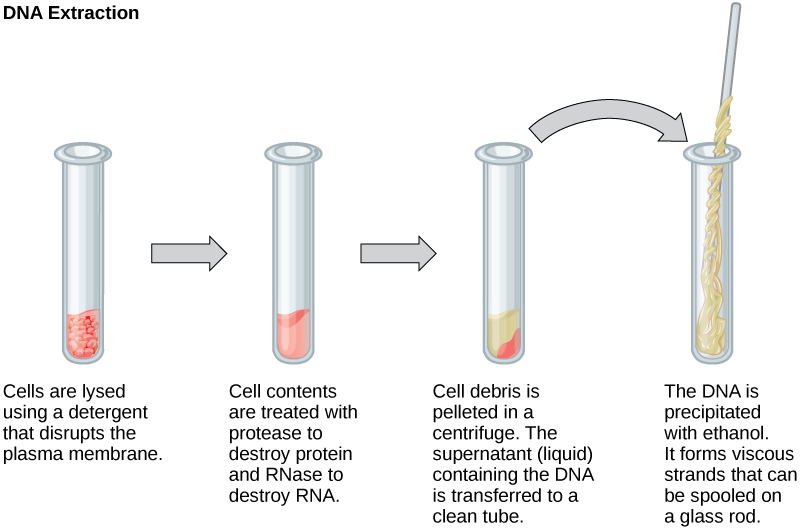
RNA is studied to empathise gene expression patterns in cells. RNA is naturally very unstable because enzymes that pause down RNA are usually present in nature. Some are even secreted past our own skin and are very hard to inactivate. Similar to Deoxyribonucleic acid extraction, RNA extraction involves the use of various buffers and enzymes to inactivate other macromolecules and preserve only the RNA.
Gel Electrophoresis
Because nucleic acids are negatively charged ions at neutral or alkaline pH in an aqueous environs, they can exist moved by an electric field. Gel electrophoresis is a technique used to split charged molecules on the basis of size and charge. The nucleic acids can be separated equally whole chromosomes or equally fragments. The nucleic acids are loaded into a slot at one end of a gel matrix, an electric electric current is applied, and negatively charged molecules are pulled toward the contrary terminate of the gel (the terminate with the positive electrode). Smaller molecules movement through the pores in the gel faster than larger molecules; this difference in the charge per unit of migration separates the fragments on the basis of size. The nucleic acids in a gel matrix are invisible until they are stained with a compound that allows them to be seen, such as a dye. Distinct fragments of nucleic acids appear every bit bands at specific distances from the top of the gel (the negative electrode end) that are based on their size (Figure ten.3). A mixture of many fragments of varying sizes announced as a long smear, whereas uncut genomic DNA is usually too large to run through the gel and forms a unmarried large band at the top of the gel.
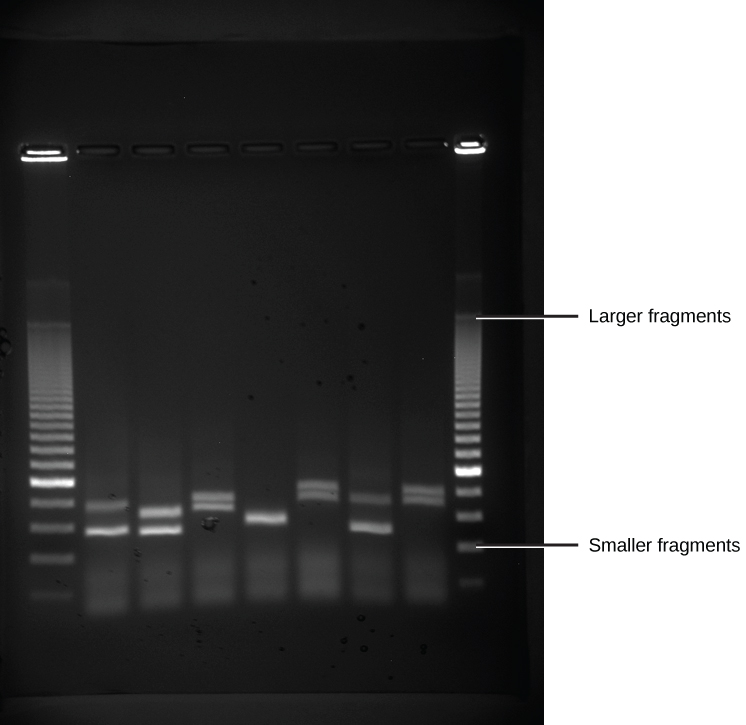
Polymerase Chain Reaction
Deoxyribonucleic acid analysis often requires focusing on one or more specific regions of the genome. It also frequently involves situations in which only one or a few copies of a DNA molecule are available for further assay. These amounts are insufficient for most procedures, such as gel electrophoresis. Polymerase concatenation reaction (PCR) is a technique used to rapidly increase the number of copies of specific regions of Deoxyribonucleic acid for further analyses (Figure 10.4). PCR uses a special form of DNA polymerase, the enzyme that replicates Deoxyribonucleic acid, and other short nucleotide sequences called primers that base pair to a specific portion of the Dna beingness replicated. PCR is used for many purposes in laboratories. These include: 1) the identification of the possessor of a DNA sample left at a crime scene; 2) paternity analysis; 3) the comparison of small amounts of ancient Deoxyribonucleic acid with modern organisms; and four) determining the sequence of nucleotides in a specific region.
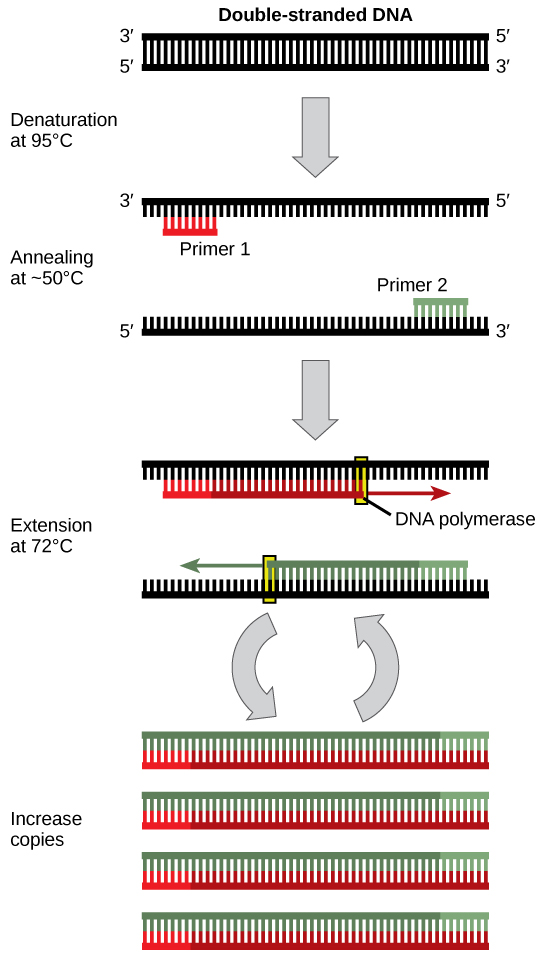
Cloning
In full general, cloning means the creation of a perfect replica. Typically, the word is used to draw the creation of a genetically identical copy. In biology, the re-creation of a whole organism is referred to as "reproductive cloning." Long before attempts were made to clone an entire organism, researchers learned how to copy curt stretches of DNA—a process that is referred to as molecular cloning.
Molecular Cloning
Cloning allows for the creation of multiple copies of genes, expression of genes, and written report of specific genes. To get the DNA fragment into a bacterial prison cell in a form that will exist copied or expressed, the fragment is first inserted into a plasmid. A plasmid (also called a vector in this context) is a small circular Deoxyribonucleic acid molecule that replicates independently of the chromosomal DNA in bacteria. In cloning, the plasmid molecules tin exist used to provide a "vehicle" in which to insert a desired Dna fragment. Modified plasmids are usually reintroduced into a bacterial host for replication. As the bacteria divide, they copy their ain Dna (including the plasmids). The inserted Dna fragment is copied forth with the rest of the bacterial Deoxyribonucleic acid. In a bacterial cell, the fragment of DNA from the human genome (or some other organism that is existence studied) is referred to equally strange DNA to differentiate information technology from the DNA of the bacterium (the host Deoxyribonucleic acid).
Plasmids occur naturally in bacterial populations (such as Escherichia coli) and have genes that tin can contribute favorable traits to the organism, such as antibiotic resistance (the ability to be unaffected by antibiotics). Plasmids take been highly engineered as vectors for molecular cloning and for the subsequent large-calibration production of important molecules, such equally insulin. A valuable characteristic of plasmid vectors is the ease with which a foreign DNA fragment can be introduced. These plasmid vectors contain many short Deoxyribonucleic acid sequences that can be cut with unlike usually bachelor restriction enzymes. Restriction enzymes (besides called restriction endonucleases) recognize specific DNA sequences and cut them in a predictable manner; they are naturally produced by bacteria equally a defence machinery confronting foreign DNA. Many brake enzymes make staggered cuts in the two strands of Dna, such that the cut ends accept a 2- to 4-nucleotide single-stranded overhang. The sequence that is recognized by the restriction enzyme is a iv- to eight-nucleotide sequence that is a palindrome. Like with a word palindrome, this means the sequence reads the same frontwards and backward. In almost cases, the sequence reads the same forward on one strand and astern on the complementary strand. When a staggered cut is made in a sequence like this, the overhangs are complementary (Figure 10.5).
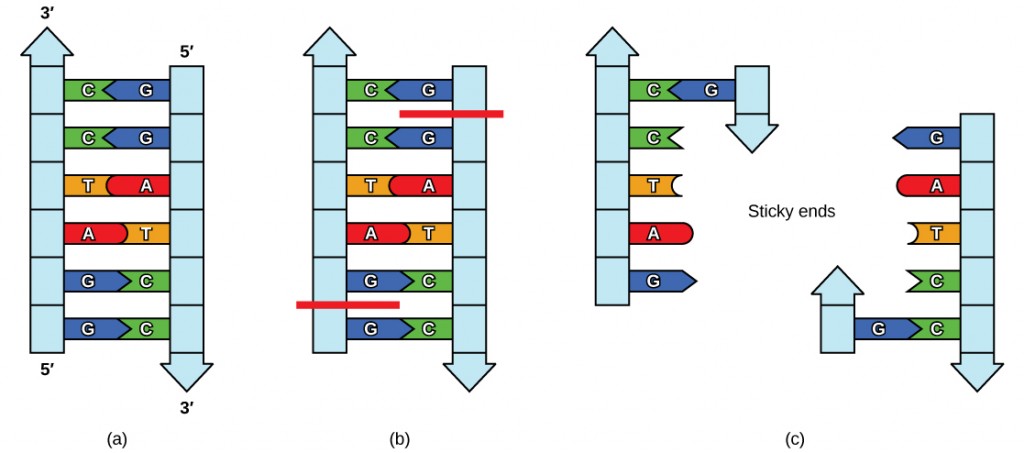
Because these overhangs are capable of coming back together by hydrogen bonding with complementary overhangs on a piece of Dna cutting with the aforementioned restriction enzyme, these are called "viscous ends." The procedure of forming hydrogen bonds between complementary sequences on unmarried strands to form double-stranded DNA is called annealing. Addition of an enzyme called Dna ligase, which takes function in Deoxyribonucleic acid replication in cells, permanently joins the Deoxyribonucleic acid fragments when the gummy ends come together. In this way, any DNA fragment can be spliced between the two ends of a plasmid DNA that has been cut with the same restriction enzyme (Figure 10.6).
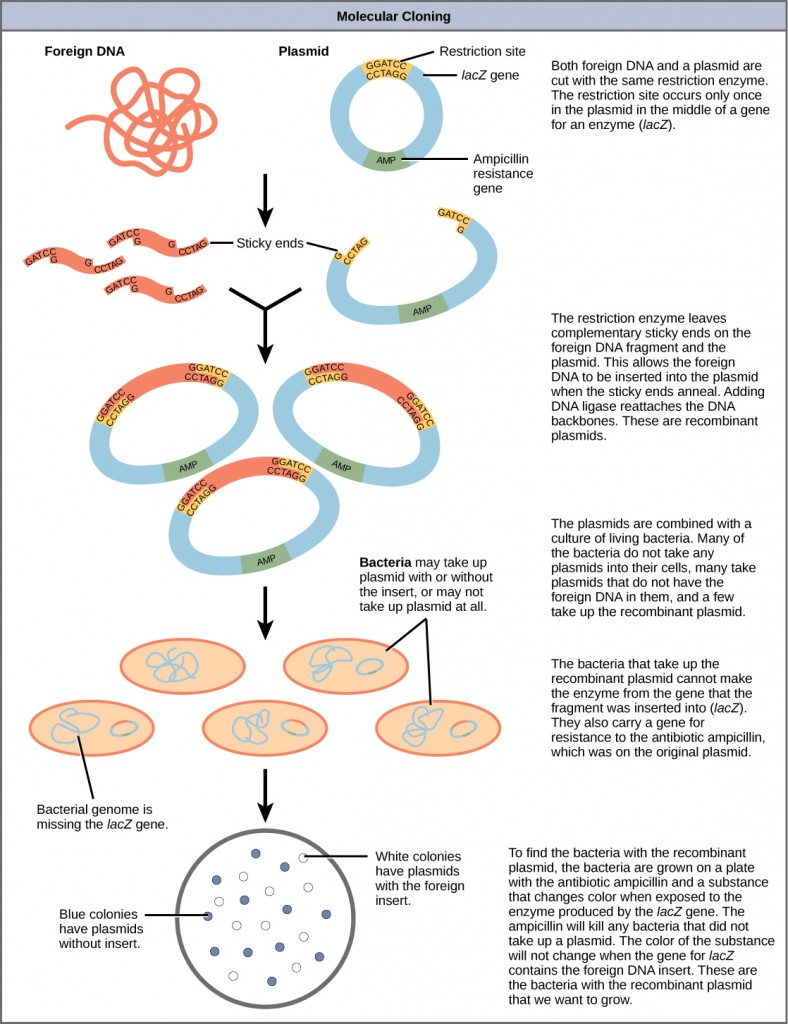
Plasmids with foreign Dna inserted into them are chosen recombinant Deoxyribonucleic acid molecules considering they incorporate new combinations of genetic cloth. Proteins that are produced from recombinant DNA molecules are called recombinant proteins. Non all recombinant plasmids are capable of expressing genes. Plasmids may also be engineered to express proteins only when stimulated by certain environmental factors, and so that scientists can control the expression of the recombinant proteins.
Reproductive Cloning
Reproductive cloning is a method used to make a clone or an identical re-create of an entire multicellular organism. Well-nigh multicellular organisms undergo reproduction by sexual ways, which involves the contribution of DNA from two individuals (parents), making it impossible to generate an identical copy or a clone of either parent. Contempo advances in biotechnology take made it possible to reproductively clone mammals in the laboratory.
Natural sexual reproduction involves the union, during fertilization, of a sperm and an egg. Each of these gametes is haploid, meaning they contain one prepare of chromosomes in their nuclei. The resulting cell, or zygote, is and then diploid and contains two sets of chromosomes. This cell divides mitotically to produce a multicellular organism. However, the union of only any two cells cannot produce a feasible zygote; there are components in the cytoplasm of the egg cell that are essential for the early development of the embryo during its first few cell divisions. Without these provisions, there would exist no subsequent evolution. Therefore, to produce a new individual, both a diploid genetic complement and an egg cytoplasm are required. The arroyo to producing an artificially cloned individual is to take the egg prison cell of ane individual and to remove the haploid nucleus. Then a diploid nucleus from a torso cell of a second individual, the donor, is put into the egg jail cell. The egg is so stimulated to divide so that development proceeds. This sounds simple, only in fact information technology takes many attempts before each of the steps is completed successfully.
The first cloned agricultural creature was Dolly, a sheep who was born in 1996. The success charge per unit of reproductive cloning at the fourth dimension was very low. Dolly lived for 6 years and died of a lung tumor (Effigy 10.seven). There was speculation that because the cell Deoxyribonucleic acid that gave rise to Dolly came from an older private, the historic period of the DNA may have affected her life expectancy. Since Dolly, several species of animals (such as horses, bulls, and goats) accept been successfully cloned.
There have been attempts at producing cloned human being embryos as sources of embryonic stem cells. In the procedure, the Deoxyribonucleic acid from an developed human is introduced into a human egg prison cell, which is then stimulated to divide. The technology is like to the technology that was used to produce Dolly, only the embryo is never implanted into a surrogate mother. The cells produced are chosen embryonic stem cells because they have the capacity to develop into many dissimilar kinds of cells, such equally muscle or nerve cells. The stem cells could be used to research and ultimately provide therapeutic applications, such as replacing damaged tissues. The benefit of cloning in this instance is that the cells used to regenerate new tissues would be a perfect lucifer to the donor of the original DNA. For example, a leukemia patient would not require a sibling with a tissue friction match for a bone-marrow transplant.
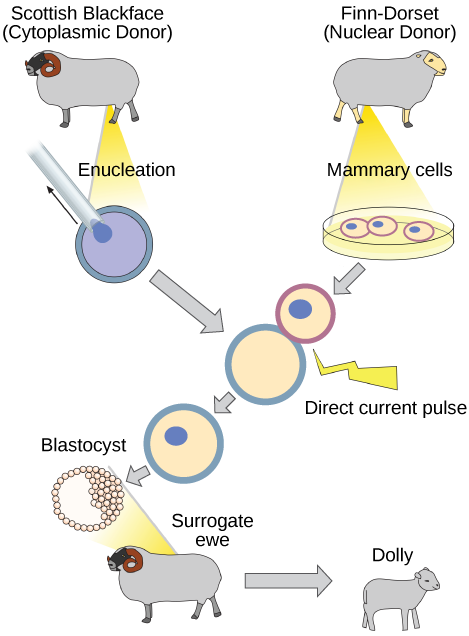
Why was Dolly a Finn-Dorset and non a Scottish Blackface sheep?
Considering even though the original cell came from a Scottish Blackface sheep and the surrogate mother was a Scottish Blackface, the DNA came from a Finn-Dorset.
Genetic Engineering
Using recombinant DNA engineering to alter an organism's Dna to achieve desirable traits is called genetic engineering. Improver of foreign Deoxyribonucleic acid in the grade of recombinant Deoxyribonucleic acid vectors that are generated by molecular cloning is the most mutual method of genetic engineering science. An organism that receives the recombinant DNA is called a genetically modified organism (GMO). If the foreign Dna that is introduced comes from a dissimilar species, the host organism is chosen transgenic. Bacteria, plants, and animals accept been genetically modified since the early 1970s for academic, medical, agricultural, and industrial purposes. These applications will be examined in more detail in the next module.
Concept in Activity

Watch this short video explaining how scientists create a transgenic creature.
Although the classic methods of studying the office of genes began with a given phenotype and adamant the genetic footing of that phenotype, modernistic techniques allow researchers to outset at the DNA sequence level and enquire: "What does this gene or DNA element do?" This technique, called reverse genetics, has resulted in reversing the classical genetic methodology. One example of this method is analogous to damaging a body role to decide its part. An insect that loses a wing cannot wing, which means that the fly's function is flight. The classic genetic method compares insects that cannot fly with insects that can wing, and observes that the non-flying insects have lost wings. Similarly in a opposite genetics approach, mutating or deleting genes provides researchers with clues about cistron role. Alternately, contrary genetics can be used to cause a gene to overexpress itself to determine what phenotypic effects may occur.
Section Summary
Nucleic acids can be isolated from cells for the purposes of further analysis past breaking open the cells and enzymatically destroying all other major macromolecules. Fragmented or whole chromosomes tin be separated on the basis of size by gel electrophoresis. Brusque stretches of Deoxyribonucleic acid can exist amplified by PCR. DNA can be cut (and subsequently re-spliced together) using restriction enzymes. The molecular and cellular techniques of biotechnology permit researchers to genetically engineer organisms, modifying them to accomplish desirable traits.
Cloning may involve cloning pocket-sized DNA fragments (molecular cloning), or cloning unabridged organisms (reproductive cloning). In molecular cloning with bacteria, a desired Deoxyribonucleic acid fragment is inserted into a bacterial plasmid using restriction enzymes and the plasmid is taken upward past a bacterium, which will then limited the strange Dna. Using other techniques, foreign genes can be inserted into eukaryotic organisms. In each case, the organisms are called transgenic organisms. In reproductive cloning, a donor nucleus is put into an enucleated egg prison cell, which is then stimulated to divide and develop into an organism.
In opposite genetics methods, a gene is mutated or removed in some way to identify its effect on the phenotype of the whole organism every bit a mode to make up one's mind its part.
Glossary
anneal: in molecular biology, the process by which two single strands of Dna hydrogen bond at complementary nucleotides to form a double-stranded molecule
biotechnology: the employ of artificial methods to modify the genetic material of living organisms or cells to produce novel compounds or to perform new functions
cloning: the production of an exact copy—specifically, an exact genetic copy—of a gene, cell, or organism
gel electrophoresis: a technique used to separate molecules on the basis of their ability to migrate through a semisolid gel in response to an electric electric current
genetic engineering science: alteration of the genetic makeup of an organism using the molecular methods of biotechnology
genetically modified organism (GMO): an organism whose genome has been artificially changed
plasmid: a small circular molecule of DNA constitute in bacteria that replicates independently of the main bacterial chromosome; plasmids lawmaking for some important traits for bacteria and can exist used as vectors to transport Deoxyribonucleic acid into bacteria in genetic engineering science applications
polymerase concatenation reaction (PCR): a technique used to make multiple copies of Dna
recombinant Dna: a combination of Deoxyribonucleic acid fragments generated by molecular cloning that does not exist in nature
strong>recombinant protein: a poly peptide that is expressed from recombinant DNA molecules
restriction enzyme: an enzyme that recognizes a specific nucleotide sequence in DNA and cuts the Dna double strand at that recognition site, often with a staggered cutting leaving brusque single strands or "sticky" ends
reverse genetics: a course of genetic analysis that manipulates Deoxyribonucleic acid to disrupt or affect the production of a gene to clarify the gene's function
reproductive cloning: cloning of entire organisms
transgenic: describing an organism that receives Deoxyribonucleic acid from a different species
Source: https://opentextbc.ca/biology/chapter/10-1-cloning-and-genetic-engineering/
Posted by: alcarazliplet.blogspot.com

0 Response to "Is An Organism's Genetic Makeup Working Behind The Scenes"
Post a Comment